Those of us working on an alternative history of humanity need to hold ourselves to standards of evidence AT LEAST AS HIGH as is demanded of mainstream scholars if we are ever to get history rewritten. With this in mind I want to say a few words about the “ancient astronaut hypothesis” and its claim that revelations and radical new meanings emerge from ancient texts and traditions when they are read in the light of modern space-exploration technology.
I have no problem with the notion that the universe is full of life, or with the notion that we are being visited now, or have been visited in the past, by “astronauts” from other planets. Much remains unknown and while that is the case we should not be too hasty to ignore extraordinary possibilities.
I am quite clear, however, having spent more than quarter of a century walking the walk across many of the most intriguing ancient archaeological sites on earth, and digging into ancient texts and traditions from all around the world, that NO ancient archaeological site and NO ancient text or tradition that I have yet come across provides persuasive evidence for the “ancient astronaut hypothesis”.
And I’m equally clear that in order to give the impression that the sites and texts do support their hypothesis, it is necessary for ancient astronaut theorists again and again to take the ancient texts out of context, or to misquote them deliberately, rather than to present them to readers in an honest and transparent way.
Please DON’T send me screeds of the late Zecharia Sitchin’s “translations” by way of refutation of this! If you were to spend as much time as I have examining the original source material upon which Sitchin drew, and comparing what you find to how Sitchin presented it in his Earth Chronicles series, you would, I think, be horrified by the scale of the illusion that has been created: a gigantic work of science fiction masquerading as fact, and giving rise to something like a New Age “religion” in which people have “faith” in the existence of a planet called Nibiru and its advanced, high-tech inhabitants — our creators, no less! — known as the Nefilim or the Anunnaki.
I am in the midst of writing a lengthy article on this subject which I will post here in due course. Meanwhile I have a question. Is anyone reading this aware of an author or advocate of the “ancient astronaut” hypothesis who documents any kind of technological PROGRESS amongst our supposed alien overlords? For example, in the case of Zecharia Sitchin’s work, we are told that the Nefilim/Anunnaki first visited Earth 450,000 years ago and that they thereafter made repeated visits over many millennia, during which long span of time evidence begins to emerge of “Mankind’s march to civilization through the intervention of the Nefilim”. In particular Sitchin claims that crucial interventions occurred around 11,000 BC, around 7400 BC and around 3800 BC – dates that are each separated from one another by intervals of 3600 years, the period of the alleged orbit of Nibiru. (The 12th Planet, 2007 paperback edition, p. 251).
As readers may be aware Sitchin first published The 12th Planet in 1976 – at a time when NASA had been aggressively exploring near-Earth space for about 13 years. I find it curious, therefore, that Sitchin’s Nefilim allegedly used space-rocket technology identical to that deployed by NASA in the 1960’s and 70’s. My question is this — at any point in the Sitchin opus has any reader come across reports of any enhancements and improvements being implemented in “Nefilim” spaceships – or are we to believe that they remained for hundreds of thousands of years at the technological level we ourselves had reached at around time Sitchin wrote The 12th Planet and subsequent volumes of the Earth Chronicles series — namely of “fiery rockets”, space shuttles with long glide paths and colossal “landing platforms” to take the weight of the alleged rockets/shuttles”?
Did Sitchin FIND evidence of alien space tech in the cuneiform texts, or did he PROJECT late 20th century NASA technology onto his imagined ancient astronauts?
My own view is that all of the anomalies of history and prehistory pointed to by advocates of the ancient astronaut hypothesis are far better and more elegantly explained as emanating from a lost, advanced HUMAN civilization of prehistoric antiquity than from high-tech alien visitors from another planet. I’ll be making the case for this at Contact in the Desert, Joshua Tree, California, 3-6 June. As well as my lecture on this subject to the full conference I’ll be hosting a workshop on the “consciousness connection” to ET abduction and encounter experiences — which I think are evidence of something MUCH more mysterious going on than JUST contacts with physical beings a bit like us but from another planet and with higher tech. Finally I will also be holding an open-house “Intensive” for discussion with critics and supporters of my work. Let’s discuss everything! Details of my main presentation, and for registration at my workshop and intensive can be found by clicking this link: http://contactinthedesert.com/grahamhancock/. Hope to see you in the desert!!
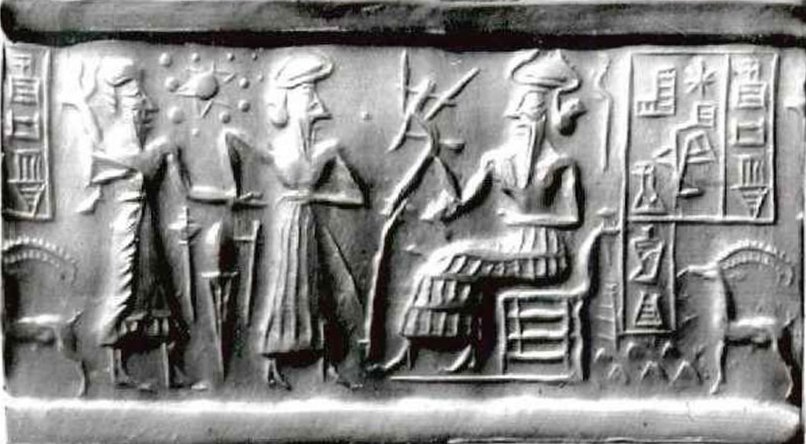
The illustration accompanying this post shows Mesopotamian seal VA 243, kept in the Vorderasiatische Museum in Berlin. This seal is the centerpiece of Zecharia Sitchin’s theory that the Sumerians had advanced astronomical knowledge of the planetary bodies in our solar system. For a critique of Sitchin’s take on the seal see http://www.michaelsheiser.com/va_243%20page.htm, with more detailed information available on this pdf: http://www.sitchiniswrong.com/VA243/VA243.htm


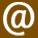




The best evidence for ancient astronauts is in the precision and or irrationally large stonework at the Great Pyramid, Puma Punku, and Baalbeck as perhaps the best examples which perplex modern day science. Since we’re not finding entire cities made from this kind of construction, only isolated examples, I think they were left as “Monoliths” ( as in 2001, A Space Odyssey ) to influence or inspire wonder among prehistoric humans and challenge them to “figure it out”. The machinery required to make them would have to have been equally large and impressive. So where are the machines? Like any construction site, you wouldn’t leave your cranes, backhoe’s, or table saws around when the jobs done. Of course not, you take them with you. Whoever built them didn’t stay, otherwise we would be finding advanced mega cities the size of London, New York, or Tokyo.
Two words, Kreg Hines: “Gobekli Tepi”. An entire complex of megalithic construction, and not even “an isolated example”.
Do your homework! Read something. Check the references. Don’t just throw out idle comments and imaginings; doing so only muddies the waters and sets back the cause of alternative history.
Though Gobeckli Tepi is indeed a groundbreaking megalithic site, it doesn’t exhibit precise machining of 1/100th of an inch that is in evidence at Giza or Puma Punku. Just as in Stonehenge, he stones are large and arranged to match astrological alignments but could have been achieved by human labor alone, so I’m differentiating between megalithic and advanced megalithic. The work of Chris Dunn and Brien Forester clearly demonstrate the best evidence of stonework that can only be achieved by computerized machining. Certainly there are many examples of megalithic sites that are more widespread around the world, but few exhibit advanced machining in their construct.
Considering the time scales involved, (4000+ years at minimum, possibly 10000+ in the case of the Sphinx, Gobekli Tepe, or Puma Punku) what sort of building machines would be left? Factor in multiple global catastrophes of the cometary impact type to “wipe the slate clean”, not to mention people’s unfortunate tendency to destroy things they don’t understand, or “believe in”, whatever THAT means, (Arabs stripping the Great Pyramid casing stones, Spanish conquistadors destroying temples and “instruments of magic/the devil” [technology?], etc.) and we are left with what we have: giant blocks of stone with no reliable record of who built them. Consider also the value of advanced materials like metal, glass, crystal, etc., especially after global catastrophes that probably destroyed the true knowledge of the society that came before. Check out the History Channel doc “Life After People” for an idea of what would be left of US after such a catastrophe. To paraphrase Joe Rogan: If you were left in the jungle with just a knife, how long before you could send an e-mail?
Ancient aliens is total crap written by stupid conspiracy brainwashed ufo americans
In the well known picture shown at the top of the page…. I have many so called experts talk about the solar system (shown between the two standing men) They all point and identify the different planets and moons…..
How come no one ever talks or refers to the lone planet or moon shown directly in front of the point of the middle mans nose???? Could this be planet X way out on its elliptical orbit…
If not Which Planet or Moon does it represent???
Thank you for the clarity!
Have you ever checked out the stone ruins of South Africa? Michael Tellinger has brought these to the front of my thought. Although he clearly loves the “Ancient Astronaut Hypothesis” there are these stone structures there. They do look like cymatic patterns when viewed from above. They are interconnected and feature a strange resonance.
I was curious if you have given much thought to these formations?
Thank you,
http://michaeltellinger.com/ancient-civilisations/
After studying the majority of what both you and the Ancient Astronaut Hypothesis have written, I tend to believe you guys need to merge and work together as I think you are both right. Extraterrestrial interaction with the unintended consequences, a violation or ignorance of the “prime directive in Star Trek” led humans out of the caves and away from their animal minded relatives on this planet, rather through teaching or perhaps even genetic manipulation. With this came the “Middle Men” between the ET’s and the primitive humans “Managers” if you will to organize our society much like a modern corporation handing down need to know information to the workers. Rather all of humanity had this ET contact or just some it led to the advanced civilization with technology and building styles we can’t duplicate today that you discuss in your books. Once the ET’s left or ceased constant contact for whatever reason these Managers became our royal blood lines “by having the ear of god at there disposal” and the ET’s arcane teachings that only the “Managers” knew and the mysteries surrounding them became our world religions. This explains how such great technology and information could been known so long ago by such ancient civilizations and why ET’s and UFO’s as they likely exist only seem to be interesting in a low impact reconnaissance of our planet and not out right contact.
Evidence that humans were made or genetically altered by ET’s.
http://www.ancient-code.com/scientists-have-found-an-alien-code-in-our-dna-ancient-engineers/
Evidence of ET contact in Humanities ancient past.
https://www.youtube.com/watch?v=idUgHlJ3IHY
Zecharia Sitchin’s work is a joke. How can anyone take his theories serious? He took bits and pieces from the Bible and other ancient texts and didn’t really understand what they said. And presto! There was his “evidence” for “astronauts” from ancient times. If we humans were pushed forward by some “extraterrestrials”, I’m convinced they were not astronauts from a far away planet. But since the modern man has cut out all thoughts of spirituality from the equation, maybe we never will find out the real truth of our past.
Thank you Graham for spelling out your position, in the paradigmatic spectrum of popular science.
Here is my response;
http://grahamhancock.com/phorum/read.php?1,1051077,1051077#msg-1051077
India is a living example of ancient civilization. which everyone seem to neglect and they try to find ancient civilizations from ruined instead of from a living one amazing you Western scholars. Who are on a destructive way to destroy or modify and Christianize ancient knowledge. who are trying to bring this world to dark age and destruction of human beings and destruction of earth. Before this idiots change every knowledge and everything will be gone.
Hi,
it seems to me that the “ancient astronaut”camp has grown because of the publicity, and not because of rational thought.
We have a situation where we have many ancient artifacts that have been found in an era when the technology used to make them was not available on earth, therefore, aliens must have made them.
Given that we have since found that the sea level has changed radically since these artifacts were deposited, that the towns and cities along ancient coast lines have been inundated, and that any civilization retreating in the face of ever rising flood waters would deposit its’ knowledge with their engineers and other technologists, who, over time, would become the keepers of knowledge, and eventually the priesthood, is it not possible that the original theories of how they were made and what their purpose was are not acurate?
To accept that a person who found an artifact 150 years ago, and ascribed a purpose to it based on his knowledge of the civilization that he thinks made it, might have been wrong should be a natural and accepted process, but from what I have read, anyone who reviews the works of such people and comes up with another answer is immediately reviled. Such are the workings of self-interested acedemia.
They stated that any large building HAD to be a place of worship,or burial, because that was the only use that “primitive” people could possibly have had for such a place……. yeah, right.
For the current acedemics to state that the ancient stonework examinde by Christopher Dunn, and found to be consistently accurate, was made using primitive, and as we now know,unsuitable tooling shows that they are stuck in their ways, and thoughtprocesses, as well as being frightened of losing their tenures.
So much of the derision hurled at people such as Mr Hancock by the media and related Western acedemia is based on what they regard as unacceptable timescales.
Why is this, when the archeologists in the Eastern world have no problems with these timescales, due to the timescales used by the ancient Hindu peoples and others of their like?.
Too many vested interests, with too much money and too many reputations at stake would seem to be a reasonable reply.
Keep at it Graham, something undeniable will turn up one day, and as long as an acredited Archeologist, with an independent film crew or journalist, who doesn’t work for a major media outlet that can be “persuaded” to bury it, is present, the world could change
Where the Egyptians speak of a First Time, and Buddhists refer to the “first time knowledge was imparted to humanity by a Buddha”, the Dogon speak of a time when “humanity was restored to culture”, clearly implying a belief that humanity had previously attained a degree of high culture. Meanwhile, both the Buddhists and Dogon explicitly claim instructional contact in ancient times from a non-human source. Likewise, there is a specific and sensible rationale, based on Dogon and Buddhist claims, for at least entertaining the idea of non-human interaction. The significant upshot of the set of claims is that we may be considering a false dichotomy. For me, it’s thinkable that both viewpoints may have a basis in truth.
From my understanding, there was a time when all mankind knew how to attain enlightenment; knowledge lost at the time of the flood. Could it be that spiritual enlightenment was perceived as coming to us from a non-human and therefore external source of wisdom because of its very nature? Is that what these ancient cultures were referring to? I agree with Laird Scranton; that would neither obviate the ancient alien theory nor do anything for it.
The evidence is abundant but its always surpressed by low vibrational thinking and a European ruling elite agenda to control anything that is beneficial to the masses tellingerand others find substantial amounts of evidence as sincere as they may be lack a fully functional pineal gland to look holistically into what’s in front of them the pineal gland is the connection to the cosmos which is the essence of all that is not understood regarding ancient technology
Nearest star system closest to our sun is 4.22 Light Years away.
Assuming the fastest space ship with a speed of 100,000 mph; the time to travel to Alpha/Proxima Centauri: speed of light; 186,000 mps so, 186,000 x 3600 x 24 x 365 x 4.22 =
24,753,237,120,000 miles.
At 100,000 mph this would merely take 247,532,371.2 hrs, or 28,257.12 years.
While it is estimated that there might be as many as 40 Billion habitable planets within our sight, the nearest one is at least 14 light years away, or 3 times the distance.
That means give or take 100,000 years.
Have ancient civilisations come here?
They would have been sorely disappointed finding Homo Erectus as the most intelligent species on our planet.
If they had the capability of that kind of travel, they would find even us not much of an improvement.
well said.
Our sense of what’s reasonable or possible changes with knowledge. There were eras in which the conventional wisdom was that humans couldn’t withstand travel speeds of 30 MPH.
I really think graham that despite your celeb status on this topic, you are discounting ancient texts completely!I assume you have seen the LAH expeditions and their conclusions?! I really can’t understand why u are debunking this so overtly?, for someone who’s been on countless Ancient Aliens episodes this looks like your deliberate trying to squash this theory, I wish sitchin was still alive to defend his corner, u obviously have no idea the lengths he went to for this theory. Why not attack von daniken?! His book was out first. Or is it that he is still alive to fight his corner? Discounting the ancient texts as myth the way you seem to be doing isn’t professional. One things for sure, this will defo help the sales of Magicians of the gods! Which btw is a great book.
I’ll take humans, in pre or present form over any ancient alien hypo any day!
While I think that you Mr. Hancock are fundamentally correct, I do think that our ancient ancestors were also influenced by other intelligences at key points in time. Whether or not they were extraterrestrials who traveled threw other dimensions to reach the physical Earth, or they originated from other dimensions, is the question. Some may have been one way and some another. These beings from other dimensions would be the Gods of the ancients. The extraterrestrials would have also been an influence on the ancients. Of course I agree with you that our mortal ancestors were far more advanced materially, mentally, and spiritually than they are usually given credit for. So mankind’s lost history was actually created by the dynamic intercourse of these three groups. That is the conclusion I have come to anyway. Your thoughts on my theory?
I would like to add that I agree with you about Sitchin; at best he was guilty of very sloppy scholarship and at worst he was an outright fraud. While I don’t agree entirely with Von Daniken, he does make some valid points in certain specific instances by way of contrast.
I am a great admirer of yours and I truly appreciate all of your fine work. You have taught me a great deal. Keep up the fine work.
I don’t think it is necessary to introduce alien contact to explain our intelligence, and scientifically, the preferred theory is the simplest. Since we are the same species across the world – from Australian Aborigines to uncontacted Amazonians, it must follow that our original exodus from Africa was when we were as we are now: there cannot have been identical evolution over the spread of the earth. If we haven’t therefore evolved since then, it follows that we were just as intelligent as we are now – and current thinking is that this was 80,000 years ago. So plenty of time to have had civilisations rise and fall. How long did the Roman Empire last? Back to our ancient civilisation, what about the idea put forward yesterday that the whole of Stonehenge was moved from Wales? Does this give greater credence to the idea that the megalithic builders were seafarers who first inhabited the west coast? The Gaelic connections from Brittany to Wales and Scotland make more sense as an arrival point, than that they were pushed out from the east.
The question is, when ancient cultures specifically claim non-human contact,on what grounds do we dismiss it without first exploring the reasonableness of the claim.
If they did visit it would be at different consciouness levels. Not dense material like sticthen proposes.
They would be immaterial that’s why there vehicles ufos disappear interdimensional maybe. Who knows just get sick of all the mass disinformation. But I sure know sticthen works are bogus, just gives us a different platform to work with that’s all, empty our cups and start again.
There is no doubt that a GLOBAL advanced human race existed 12,000-15,000 years ago that may or may not have been influenced by intelligent E.T.’s. What Graham is suggesting is mainstream science will continue to easily keep this knowledge of ice age era advanced human civilization silenced if it associated and attributed wild theories of E.T. influence on mankind. Baby steps… Get mainstream science to first accept the Human aspect, the timeline, the global connection, and level of advancement, then we get the history books changed. This will create great interest to know more among science and the general public, which will lead to more serious archeological research into sites ignored today. Then the E.T. Hypothesis may prove its self through tangible evidence found in ancient underwater sites. Sitchin was a nut, to assume intelligent life developed first on a planet with a 3600 year orbit that spends most time in the deep freeze at the edge of the solar system… If anything, Nibiru is a space vessel with an expedition that cycles past Earth. To say E.T. is incapable of reaching earth because of time it takes to travel between the stars shows ones ignorance. Science today has come up with no less than 4 propulsion methods that could make interstellar travel very possible, yet debunkers believe E.T. only has 20th century human rocket tech.
Keep up the good work Graham. We need more extensive research under 100-300 feet of water along coastlines where land based ancient civilizations are found.
Beliefs in visitors from space can be best uderstood in terms of religious thinking. Humans are innately confused and afraid, and we have a deep desire to have everything explained to us. Our highly elastic imaginations, brains wired for mythological story-telling, and ability to justify and believe anything we put our minds to, enhance our propensity to religious thinking.
at 99% Sitchin could be wrong on Nibiru, but if we look at every ancient religious text of past civilizations we could find that everyone of them talks about “Gods” that created life on Earth coming from the stars, or coming from the “doors of the sky”, or other words to describe something not from this planet..There are some evidences even on the Book of Genesis (Holy Bible).. I mean, I believe there is a common line that can link all this facts.. Maybe we received some kind of technology even before the known hystory, then we could have destroyed our society ourself and started again from scratch.. Or maybe we are “survivors” from a mass disaster (the Great Flood?) and we lost all our knowledge.. Who knows? Everything could be everything until we’ll progress with research..we need more evidences to rewrite history books.. so, keep working on
“a gigantic work of science fiction masquerading as fact”
—
Mr. Hancock, the same could be said about some parts of your own work. I enjoy watching you documents, it is like a more intelligent alternative to Hollywood science fiction entertainment, but I remain sceptical and unconvinced concerning some of your claims, specifically the connection between star constellations and the layout of some of the ancient temples and monuments such as the Egypt pyramids or Angor Wat.
1) star constellations are culture specific, ie. the Chinese had different constellations from the Hellenistic constellations
http://www.stellarium.org/wiki/index.php/Sky_cultures
yet you seem to claim that all ancient momenuments are built according to some universal constellation culture.
2) some of your supposed links between star constellations and ancient monuments are far fetched and hard (for me impossible) to believe. You claim that the Nazca spider represents the orion constellation by puting some dots into the spider body and legs. Do you really consider such vague analogies scientific proofs? Another example is the draco constellation and the Angkor wat. Why would the builders of the Angkor wat use a draco constellation from the Hellenistic tradition? Do you even recognize the Draco constellation in the sky?
http://www.allthesky.com/constellations/draco/big.jpg
just some random dots. Each civilization can project whatever creatures/animals they wish to see in those dots.
3) some of your numerology in those ancient sites is also rather speculative. For example your claims that the number 72 is encoded in the ancient monuments because 1 degree of precession takes 71.6 years. How do you even now that the ancient builders measured angles in 360 degrees and not in radians or some other form of angular measure? This is magic/wishful thinking and not science.
I hope you won’t take my comment as an attack. I am just pointing out why I remain unconvinced about some of your claims. Otherwise I am open-minded, and I believe an ice age civilization could have existed.
Found on Michael Scheiser’s site: “So the full (rather boring) inscription of VA243 reads: “Dubsiga, Ili-illat, your/his servant.”
Nothing boring about the inscription on this tablet to my mind. I offer you a more amusing translation of the text:
DUB: tablet
SI: horn (horn of the cow, but also drinking vessel)
GA: milk, soft (GA is our Gaia, the nurturer and sacred cow)
BY THIS TABLET, A HORN OF MILK
ILI / AN: sky
IL2: to raise
LA: to hang (two verbs combining mean “to hold up”)
TO THE SKY, BE RAISED
AT: Atlantis (Plato knew of it too, but much later.)also bead (hard and round), adamantine
IR3 / ARAD: saying, proverb
SU: to sink (the horn to sink meaning “to down”)
AND, LIKE PROVERBIAL ATLANTIS, SINK!
IN ACCORDANCE WITH THIS TABLET, THAT A HORN OF MILK BE RAISED AND, LIKE PROVERBIAL ATLANTIS, SUNK!
Perhaps a method of paying for the beer with a stamp…?
Except for Atlantis, the meanings here can be verified on the electronic Pennslyvania Sumerian Dictionary.