In chapter twenty of Graham Hancock’s Supernatural, a fascinating question about the contents of “junk” DNA is brought to light. At the time of publication, researchers had discovered that only the non-coding sections of DNA had special properties of long range ordering while the coding sections did not. This long range ordering conforms to the pattern found in Zipf’s law which is also a pattern found in spoken language.
While this is a remarkable finding in itself, there are many other processes in nature that have also been found to be Zipf compliant. What makes the findings of Eugene Stanley et al (1994) so amazing is that they also used information theory as a wholly separate test of the non-coding data. Using the theory which governs modern scientific treatments of data compression, storage and communication, they also found that non-coding DNA has the properties necessary for maintaining data integrity even in error prone conditions. This necessary form of redundancy, which is expressed in terms of “Shannon Entropy,” was not found in the coding portion of the DNA, nor was it found in random characters used as a control.
The fact that these two very different tests for linguistic characteristics are found in the non-coding sections leads very strongly to the conclusion that structured and useful information is contained within the non-coding sections of DNA. This initially led Hancock to posit that non-coding DNA, which makes up the vast majority (or at least around 90%) of the human genome, might actually be a repository of knowledge which could in some way be accessible. Recent science has added a great deal to this mind-blowing hypothesis.
“Junk” DNA? What are we talking about?
 DNA is a set of molecules whose shape and configuration of charges allow them to interact with various other molecules in ways that ultimately result in building and operating the super-complex micro machines we call cells. One little section of DNA, therefore, is like some of those complex little machines full of gears that can do fascinating but specific things. Now imagine creating enough different little machines to populate a factory and that factory’s purpose was to create even more tools, gears, and machines. This is a basic description of a cell.
DNA is a set of molecules whose shape and configuration of charges allow them to interact with various other molecules in ways that ultimately result in building and operating the super-complex micro machines we call cells. One little section of DNA, therefore, is like some of those complex little machines full of gears that can do fascinating but specific things. Now imagine creating enough different little machines to populate a factory and that factory’s purpose was to create even more tools, gears, and machines. This is a basic description of a cell.
 In this analogy the configuration of atoms which make up DNA are like specially shaped multi-pole magnets which self-assemble and fit together to make up the gears of the machinery. The specific way all these gears and machines fit together and how they are arranged is crucial to making the factory actually be able to create what it does. The sequence of events dictated by the arrangement of equipment can be said to be a sort of “program” and that is why we often refer to DNA this way. The simplest pieces are all pretty much the same so it’s the arrangement that makes the difference between “pieces of metal” and “a machine.” That configuration is information.
In this analogy the configuration of atoms which make up DNA are like specially shaped multi-pole magnets which self-assemble and fit together to make up the gears of the machinery. The specific way all these gears and machines fit together and how they are arranged is crucial to making the factory actually be able to create what it does. The sequence of events dictated by the arrangement of equipment can be said to be a sort of “program” and that is why we often refer to DNA this way. The simplest pieces are all pretty much the same so it’s the arrangement that makes the difference between “pieces of metal” and “a machine.” That configuration is information.
 The problem is that when we look in at the super complex machinery of DNA, we can only really separate “machines” from “hunks of metal” by the fact that we can see certain components work together to create things. We call these sections of DNA “coding” sections because they take part in creating proteins. When we examined all we knew about DNA and its functioning, we found that only a small percentage of the DNA actually seemed to do anything at all.
The problem is that when we look in at the super complex machinery of DNA, we can only really separate “machines” from “hunks of metal” by the fact that we can see certain components work together to create things. We call these sections of DNA “coding” sections because they take part in creating proteins. When we examined all we knew about DNA and its functioning, we found that only a small percentage of the DNA actually seemed to do anything at all.
This is like looking into a factory in which over 90% of what you see is machine parts in various states of configuration and assembly that don’t actually do anything and are never used at all in the production process. It’s just hanging around. Then, when the factory fully duplicates itself, it also duplicates all the seemingly pointless junk as well. The question we have to ask is if the non-coding DNA really is “junk” or not. Nature tends to get rid of superfluous things, so what is all the extra information for?
Ups and downs in the recent history of “junk” DNA
One of the first advances of note in examining this idea is the recent finding that some of the non-coding areas of DNA previously labelled “junk” have been reported to play a crucial role in modulating gene expression. Some of the previously labeled “junk” is, therefore, actually functional. In many reports on these findings, the very notion of “Junk DNA” is strongly dissuaded, as though all of the non-coding DNA has now been shown to have some purpose. This is the farthest thing from the truth, however!
Upon closer examination, it has long been known (as early as 1951 through Barbara McClintock) that gene regulation exists within the genome and that all of these regulators are not fully known or understood. Furthermore, the idea of “junk” DNA has been held up as support of an evolutionary development model and has been a hot-button issue for creation science proponents. Much of the literature and impetus to fight the idea of “junk” DNA has come through creation science sources with a specific axe to grind.
 Before the completion of the human genome project, most arguments against “junk” DNA were based upon comparative genomics. It was found that upon comparing genes between species, a number of genes in non-coding areas are shared. The further apart the species are genetically, the more “highly conserved” an identical gene sequence found in both is considered to be. Some genes are considered “ultra conserved” because they are found in widely divergent species such as rats, humans, and cows. The first indication that these areas in particular may be useful in some way was when over-expression of these ultraconserved regions was linked with certain types of cancer (Calin et al 2007). This finding is a special case which runs contrary to many other findings discussed below.
Before the completion of the human genome project, most arguments against “junk” DNA were based upon comparative genomics. It was found that upon comparing genes between species, a number of genes in non-coding areas are shared. The further apart the species are genetically, the more “highly conserved” an identical gene sequence found in both is considered to be. Some genes are considered “ultra conserved” because they are found in widely divergent species such as rats, humans, and cows. The first indication that these areas in particular may be useful in some way was when over-expression of these ultraconserved regions was linked with certain types of cancer (Calin et al 2007). This finding is a special case which runs contrary to many other findings discussed below.
Around the turn of the millennium, upon completion of the human genome project, it was known that around a mere 1.5% of the DNA is actually coding. But in September 2012 this same project group called the Encyclopedia of DNA Elements Project (ENCODE) had published a series of papers in Nature which showed that up to 76% of non-coding DNA was being transcribed. This report – that around 80% of genome is biologically active – also led to a great deal of conjecture and presumption. Scientifically speaking however, these findings were not direct evidence of real or direct biological functions, but evidence of the possibility of them.
As early as 2011 however, the same project had identified that a species of bladderwort was deleting the vast majority of non-coding DNA such that only 3% of its total genome was non-coding DNA; this was in opposition to the 98.5% found in humans. This extremely complex species was functioning quite well with a miniscule number of regulatory genes.
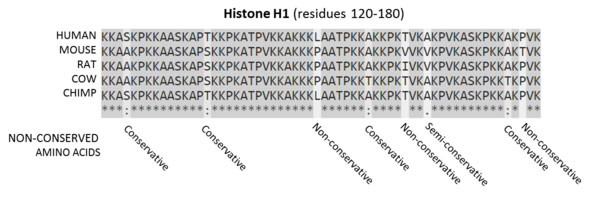 While some of the non-coding DNA in humans has been directly shown to have regulatory action, the most recent findings now show that the amount of the actual functioning non-coding DNA is still extremely small. This conclusion fits well with the bladderwort findings that only a small number of regulatory non-coding genes were necessary to function. Most notably, the ENCODE group has reported that the total functional DNA in humans is no greater than 8.2%. (Rands et al 2014) This is, however, not proof of direct functioning of even that small percentage, but an estimation made, once again, based upon comparative genomics. Only a very, very small percentage of non coding DNA has been directly shown to have regulatory action.
While some of the non-coding DNA in humans has been directly shown to have regulatory action, the most recent findings now show that the amount of the actual functioning non-coding DNA is still extremely small. This conclusion fits well with the bladderwort findings that only a small number of regulatory non-coding genes were necessary to function. Most notably, the ENCODE group has reported that the total functional DNA in humans is no greater than 8.2%. (Rands et al 2014) This is, however, not proof of direct functioning of even that small percentage, but an estimation made, once again, based upon comparative genomics. Only a very, very small percentage of non coding DNA has been directly shown to have regulatory action.
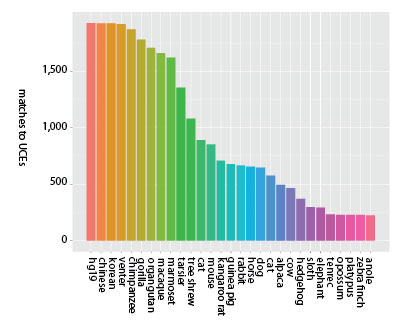 Comparative genomics in this context relies predominantly upon the conservation of genes between species, as mentioned above. It is automatically presumed that the conservation of genes occurs because of selection pressures acting upon the animal, meaning that the gene must either grant some survival advantage or otherwise play some functional role that can impact selection. Experimentation in deleting these genes, however, has shown this presumption to be highly questionable.
Comparative genomics in this context relies predominantly upon the conservation of genes between species, as mentioned above. It is automatically presumed that the conservation of genes occurs because of selection pressures acting upon the animal, meaning that the gene must either grant some survival advantage or otherwise play some functional role that can impact selection. Experimentation in deleting these genes, however, has shown this presumption to be highly questionable.
It is important to note that one of the crucial aspects of regulatory genes is that their placement near coding genes is considered the mechanism by which they function. While the first experiments which deleted highly conserved areas showed zero effect in the animals (Nobrega, M.A. et al. 2004), these extremely unexpected results were explained as perhaps related to the ease of lab conditions in keeping the animals healthy and hiding possible frailties. In another experiment however, researchers specifically chose ultra conserved areas in close proximity to genes which are known to cause severe abnormalities when not functioning normally. Once again, the deletion of these genes had absolutely zero impact upon morphology, longevity, reproduction, or metabolism (Ahituv, N. et al. 2007).
DNA data storage and information theory …again
 In the past decade and a half, the computer industry has delved deeply into the use of DNA for developing long-term, space-saving data storage. The first major proof of the concept emerged in 2012 with the writing and reading of over 5 megabits of information in DNA storage (Church et al 2012). While there have been simultaneous advancements in numerous biological computation technologies (Shaji Varghese 2015), the most telling example is the development of a DNA storage system for data which is not only nearly as easily manipulated for editing etc. as an electronic computer system, but has been shown to potentially last as long as a million years in cold storage (Grass et al 2015). Even using less than perfect data compression, all of the data on Wikipedia and Facebook combined could fit in a couple of drops of liquid and all the knowledge of mankind in the space of an elevator cabin.
In the past decade and a half, the computer industry has delved deeply into the use of DNA for developing long-term, space-saving data storage. The first major proof of the concept emerged in 2012 with the writing and reading of over 5 megabits of information in DNA storage (Church et al 2012). While there have been simultaneous advancements in numerous biological computation technologies (Shaji Varghese 2015), the most telling example is the development of a DNA storage system for data which is not only nearly as easily manipulated for editing etc. as an electronic computer system, but has been shown to potentially last as long as a million years in cold storage (Grass et al 2015). Even using less than perfect data compression, all of the data on Wikipedia and Facebook combined could fit in a couple of drops of liquid and all the knowledge of mankind in the space of an elevator cabin.
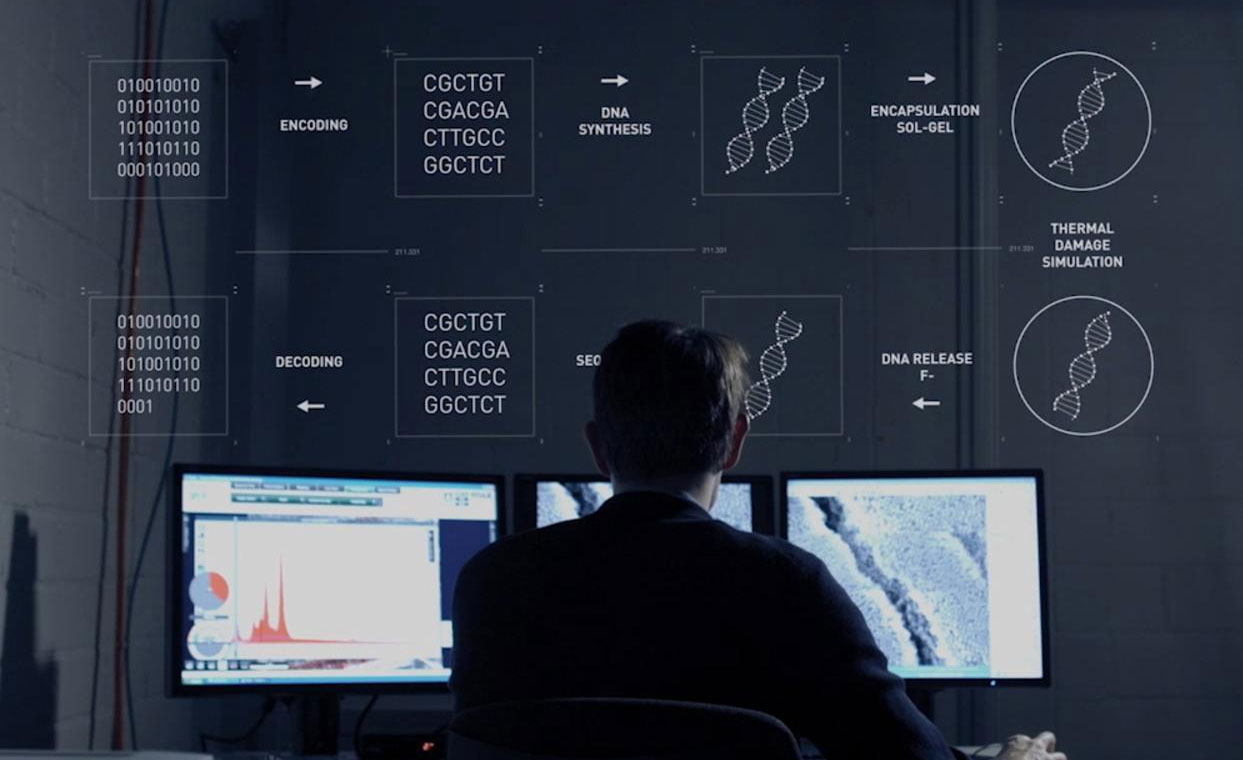
The further breakthrough in this most recent development is the application of information theory to maintain data integrity even in conditions that might cause errors. DNA can be very unstable, so the largest challenge is overcoming this limitation. By using error correction codes, lost data can be reconstructed. This is accomplished by duplicating information.
This methodology of redundancy results in precisely what was searched for (and found) in non-coding DNA in the study mentioned at the beginning of this article. With this insight, it seems as though we may have come full circle with an ancient advanced civilization who had previously discovered and used this technology.
 In the continued pursuit to design a data system which could eternally store knowledge, it is inevitable that any singular storage place for a library will be subject to destruction via simple mechanisms such as bombing, to more long term concerns such as subsumption by tectonic plates at large enough time scales. Our technology is already strongly and quickly advancing from our manipulation of DNA, towards using it as a form of wet nanotechnology (Zakeri et al 2015), which could accomplish all of the tasks that our current technologies do with the added advantage of self-assembling, self-healing and self-replicating devices.
In the continued pursuit to design a data system which could eternally store knowledge, it is inevitable that any singular storage place for a library will be subject to destruction via simple mechanisms such as bombing, to more long term concerns such as subsumption by tectonic plates at large enough time scales. Our technology is already strongly and quickly advancing from our manipulation of DNA, towards using it as a form of wet nanotechnology (Zakeri et al 2015), which could accomplish all of the tasks that our current technologies do with the added advantage of self-assembling, self-healing and self-replicating devices.
 An obvious solution to the eternal storage problem would be to write our knowledge into creatures that will grow, replicate and most importantly adapt to an ever-changing environment through branching into every possible species, to colonize every possible space and environmental niche. It makes perfect sense to do again what might have been done before.
An obvious solution to the eternal storage problem would be to write our knowledge into creatures that will grow, replicate and most importantly adapt to an ever-changing environment through branching into every possible species, to colonize every possible space and environmental niche. It makes perfect sense to do again what might have been done before.
We may decide to build an eternal library and fill it with our own books. Chances are, as we excavate the location to build this library that we will find, just under the surface, a library already built in the same way and in the same place as before.
Data means nothing without an interpretation key

One of the most crucial aspects to consider when designing this eternal library, however, is the leaving of a Rosetta stone, so that in a few million years, when a wholly different species with no concept of our language attempts to read it, it can be decoded. The Rosetta stone itself must not only be hidden in the DNA with the information, but also somehow cross language and culture barriers that are separated by an alien mental gulf of time and circumstance.
One conceivable method for creating a Rosetta stone is to cause library sections (non-coding) of the information to somehow also impact and interact with the sections of DNA which do actually form the creature (coding DNA). This could be a way of linking pure information with real world actions and objects. By combining the method of action of this “Rosetta stone” section of regulatory DNA with its location in relationship to other non-coding sections, it might represent a word within some larger statement. If this was combined with the component of the animal that a particular section of DNA represents, one might develop a seed for universal translation of the data through the symbolic actions and actors found within the DNA itself.
For instance, if the regulatory action causes a gene to “hide” or “diminish” then that sequence of DNA could be interpreted to have that meaning. If it “cuts” or “translates,” these might also give us a meaning to particular sequences. Then if it affects “blood” or an “arm” then we begin to have additional nouns to work with. All of these actions and components might be used in a metaphorical sense to build a language which can be unpacked through self-modification. Each relationship might lead to other relationships by which a well designed language might lead to ever more complex words based upon a system in which few words act as modifiers upon each other to create ever greater refinement and specificity of definition.
The discovery – that a small portion of our non-coding DNA does directly regulate some genes – is a natural conclusion to, and in further support of, the genetic library hypothesis.
Eliminating selection pressure impact on genetic information storage
 One of the other important design considerations in using animals and their offspring to store crucial information, is the ability to somehow allow the normal developmental process of species, but keep it from impacting the passing on of the data to offspring. There would need to be a selection mechanism within the genes that favors preservation of the data even though the data provides no survival advantage.
One of the other important design considerations in using animals and their offspring to store crucial information, is the ability to somehow allow the normal developmental process of species, but keep it from impacting the passing on of the data to offspring. There would need to be a selection mechanism within the genes that favors preservation of the data even though the data provides no survival advantage.
Recent development in genetics has actually provided such a mechanism and it’s called a “gene drive” or a Mutagenic Chain Reaction (MCR). In applying the CRISPR gene editing technology with a targeting sequence, researchers have created an auto-catalysing reaction that results in a self-replicating gene editing system (Gantz & Bier 2015). This system targets and edits all copies of the targeted gene and therefore not only affects the genes of a given animal, but also the genes of offspring. The editing system, passed down genetically, bypasses the normal mendelian mechanics of dominant and recessive traits by overwriting any other copy received from a normal donor parent with the gene passed down from the gene drive parent. In the experiments, the gene drive system resulted in passing down a recessive trait in fruit flies with around 97% accuracy.
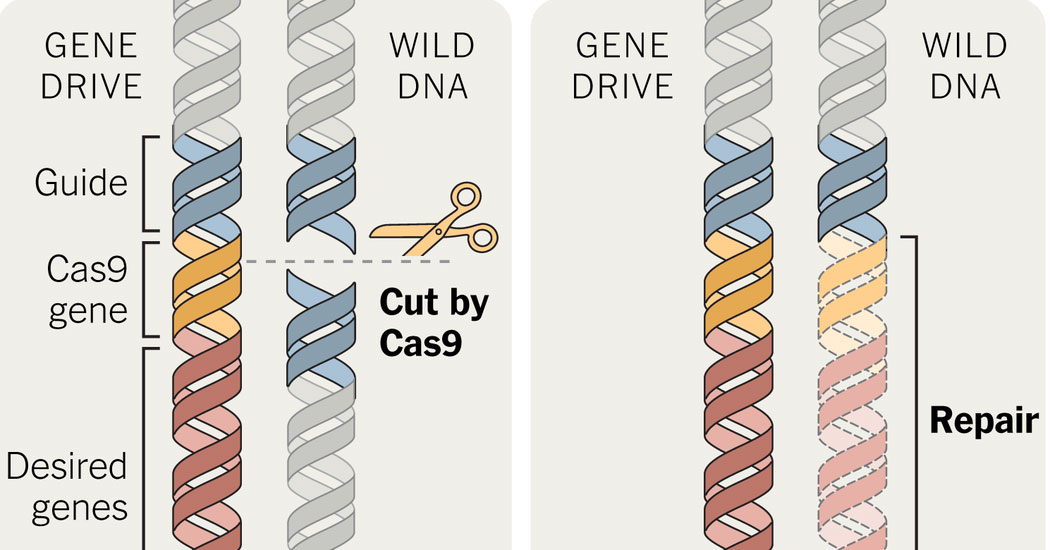 This system so completely bypasses any selection pressures that it could even result in preferentially passing down traits which directly lower the survival rate of the offspring. This system, which is entirely encapsulated within the molecular mechanics, could provide a means by which DNA-based information storage is passed down preferentially in animals completely regardless of natural selection processes.
This system so completely bypasses any selection pressures that it could even result in preferentially passing down traits which directly lower the survival rate of the offspring. This system, which is entirely encapsulated within the molecular mechanics, could provide a means by which DNA-based information storage is passed down preferentially in animals completely regardless of natural selection processes.
The end result of storing information in this way would be that after millions of years there would be ultra-conserved DNA which could be directly deleted, with zero impact on the morphology or other traits of the animals, precisely as was seen in the UCR deletion studies mentioned above.
Epigenetics, Genetic Memory, and the Library of the Ancients
Epigenetics is another crucial component to this puzzle, the knowledge of which has advanced a great deal in the past decade. We now know that the events and influences within a single animal’s life can be passed on to their offspring. Epigenetics is not a process that alters the actual DNA code but works, instead, through regulation of gene expression much like cell differentiation does. The reason we can have all the various cells of different organs appear from the single stem cell type is because of gene expression.
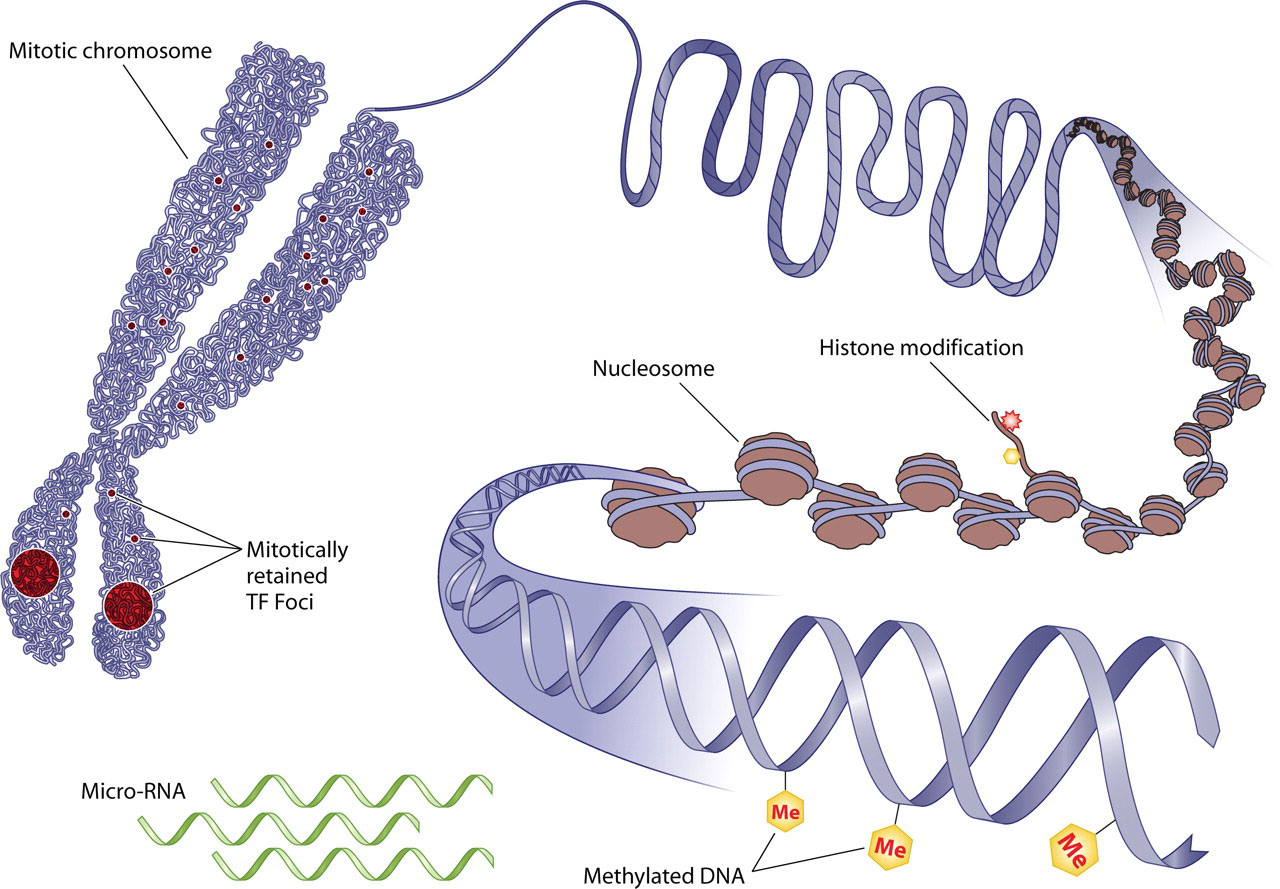
As one might expect, the sections of non-coding DNA which act in a regulatory role and impact the coding DNA have been found to be a part of the mechanism of epigenetics (Cao, J. 2014). These epigenetic changes can also, however, alter the likelihood that a real genetic mutation might occur in the affected area (Skinner 2015). The more short-term mechanisms of epigenetic change, when transmitted over longer trans-generational time scales, may, therefore, lead to direct genetic change.
The idea of genetic memory has been around for a very long time as an anecdotal conjecture. The ability of savants to know things it seems they have never learned has been explained in such a way, as have past life regressions etc. While the idea that instincts must exist as instructions hidden in genes which regulate brain morphology is widely assumed, these general instructions have generally been postulated to have formed purely through selection pressures.
With the recent understanding of epigenetics allowing for the experiences of a single individual to become heritable, the possibility of true genetic memory has become a scientifically viable area of inquiry. As such, a study was performed on mice in which parental traumatic exposure to cherry blossom scent passed down the fear conditioning to future generations in which even the grandchildren reacted very aversively toward the scent. Furthermore a gene was identified with sensitivity to that scent and changes to the expression of the gene had indeed been altered. Concurrent changes in brain structure were also observed in offspring (Dias & Ressler 2014).
‘Memories’ pass between generations – BBC

Now that genetically transmitted memory is a scientifically validated concept, it is obvious that there is a data storage mechanism in DNA which interacts with mental experiences and it seems no large presumption that the interaction could possibly be two-way in nature.
Might we then suppose that our non-coding DNA may be comprised of both trans-generational memories as well as a storehouse of more directly inserted information from long ago? Might the directly inserted older data play a role as a guiding principle for trans-generational memory and might those memories, built upon that original key, provide a sort of interpretational Rosetta stone for understanding that original information?
From a programming and engineering perspective, it makes sense to integrate the interpretational system in such a fashion.
One final thought
In 2011 another new method of identifying spoken language emerged from the application of information theory in a novel fashion. While the “Shannon Entropy” (information density) of various languages differed, one aspect remained almost exactly the same across all languages. The amount of information carried by the structure of real communicative language is higher than a word-salad mix of words. When we take any text of any language and rearrange the words randomly, the amount of information per word decreases by the same amount regardless of language (Montemurro & Zanette 2011).
My personal opinion is that if non-coding DNA were subjected to this test, it would probably fail because the language of memories is likely more symbolic than syntax based. A reliance upon word ordering is usually reserved only for spoken language and not general data storage. Therefore a failure would not, in any way, falsify the library hypothesis. However, if this test were to come back positive, not only would the library hypothesis be utterly validated, but it would also likely mean that all genetic memory is encoded directly into the language once spoken by the ancients who created this library.
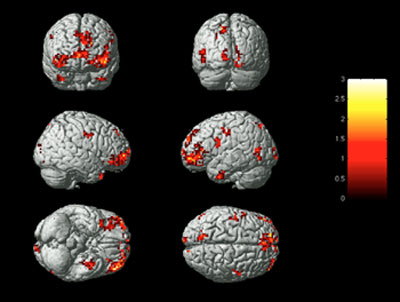
While this reliance upon syntax in the storage of genetic memory seems unlikely, it is not completely unsupported by scientific discoveries. Research at Carnegie Mellon has resulted in the discovery that language is stored so similarly from one human to the next, that the physical location in the brain for the data that represents nouns can be modeled and predicted by a computer. Tests using fMRIs to read the blood flow and activation of these areas in the brain resulted in the ability to read a subject’s mind with between 70% and 94% accuracy when the subject focused upon a noun. The accuracy was so great that this computer-modeling-based “mind-reading” technique was able to distinguish between thoughts of extremely similar objects like “pliers” and “hammer.” (Mitchell et al 2008)
While this is not a reliance upon word ordering, it shows a standardization of language storage from one human to the next. Therefore the concept of storing additional information by the relationships between words is not necessarily a large leap, but is a worthy area of future inquiry. It does, however, lend itself to a representational mechanism, based upon physical position, by which genetic memory may be encoded. Specifically, it provides a structural system by which one language may be translated into a different one.
References:
R. N. Mantegna, S. V. Buldyrev, A. L. Goldberger, S. Havlin, C.-K. Peng, M. Simons, and H. E. Stanley.
Linguistic Features of Non-coding DNA Sequences,
Phys. Rev. Lett. 73, 3169-3172 (1994)
Calin, George A. et al.
Ultraconserved Regions Encoding ncRNAs Are Altered in Human Leukemias and Carcinomas
Cancer Cell , Volume 12 , Issue 3 , 215 – 229, 2007
Ibarra-Laclette, E., Albert, V. A., Pérez-Torres, C. A., Zamudio-Hernández, F., Ortega-Estrada, M. de J., Herrera-Estrella, A., & Herrera-Estrella, L.
Transcriptomics and molecular evolutionary rate analysis of the bladderwort (Utricularia), a carnivorous plant with a minimal genome.
BMC Plant Biology (2011), 11, 101. http://doi.org/10.1186/1471-2229-11-101
Rands CM, Meader S, Ponting CP, Lunter G.
8.2% of the Human Genome Is Constrained: Variation in Rates of Turnover across Functional Element Classes in the Human Lineage.
Schierup MH, ed. PLoS Genetics. 2014;10(7):e1004525. doi:10.1371/journal.pgen.1004525.
Nobrega, M.A. et al. (2004).
Megabase deletions of gene deserts result in viable mice.
Nature 431: 988-993.
Ahituv, N. et al. (2007).
Deletion of ultraconserved elements yields viable mice.
PLoS Biology 5(9): e234.
Church, G. M., Gao, Y. & Kosuri, S.
Next-generation digital information storage in dna.
Science 337, 1628–1628 (2012)
Shaji Varghese, Johannes A. A. W. Elemans, Alan E. Rowan, Roeland J. M. Nolte,
Molecular computing: paths to chemical Turing machines,
Chem. Sci., 2015, 6, 11, 6050
Grass, R. N., Heckel, R., Puddu, M., Paunescu, D. and Stark, W. J. (2015), Robust Chemical Preservation of Digital Information on DNA in Silica with Error-Correcting Codes.
Angew. Chem. Int. Ed., 54: 2552–2555. doi:10.1002/anie.201411378
Bijan Zakeri, Timothy K Lu,
DNA nanotechnology: new adventures for an old warhorse,
Current Opinion in Chemical Biology, 2015, 28, 9
Valentino M. Gantz, Ethan Bier
The mutagenic chain reaction: A method for converting heterozygous to homozygous mutations
Science 24 Apr 2015 : 442-444
Cao, J.
The functional role of long non-coding RNAs and epigenetics.
Biological Procedures Online (2014), 16, 11. http://doi.org/10.1186/1480-9222-16-11
Skinner MK
Environmental Epigenetics and a Unified Theory of the Molecular Aspects of Evolution: A Neo-Lamarckian Concept that Facilitates Neo-Darwinian Evolution.
Genome Biol Evol. 2015 Apr 26;7(5):1296-302. doi: 10.1093/gbe/evv073.
Brian G Dias & Kerry J Ressler
Parental olfactory experience influences behavior and neural structure in subsequent generations
Nature Neuroscience 17, 89–96 (2014) doi:10.1038/nn.3594
Marcelo A. Montemurro and Damián H. Zanette.
Universal Entropy of Word Ordering Across Linguistic Families.
PLoS ONE, Vol. 6, Issue 5, May 13, 2011. DOI: 10.1371/journal.pone.0019875.
Tom M. Mitchell, Svetlana V. Shinkareva, Andrew Carlson, Kai-Min Chang, Vicente L.. Malave, Robert A. Mason, Marcel Adam Just
Predicting Human Brain Activity Associated with the Meanings of Nouns
Science, 30 MAY 2008 : 1191-1195




Having read your article and then ‘dipped into’ the article published the following day on this site I was struck by the idea that your thesis dealt with the possible structure of language used to encode knowledge in DNA where David Warner Mathisen’s article dealt with the similarity in the structure of celestial myths and metaphors. One can distil the concept of Hermetiscism into the phrase, ‘as above, so below’ and wondered whether the structures used to encode knowledge in myth and metaphor might lead to some insight as to how knowledge encoded in DNA might be arranged. Humans are very good at patterns and I suspect patterns may turn out to be key.
With all Best Wishes
Rick Standley
. . . Gives new meaning to the phrase, “Food for thought,” nearly down to the molecular constructs of that assembly of parts that yield meaning by word of mouth, multi-media, and in print.
Thank you
“Having read your article and then ‘dipped into’ the article published the following day on this site I was struck by the idea that your thesis dealt with the possible structure of language used to encode knowledge in DNA where David Warner Mathisen’s article dealt with the similarity in the structure of celestial myths and metaphors. One can distil the concept of Hermetiscism into the phrase, ‘as above, so below’ and wondered whether the structures used to encode knowledge in myth and metaphor might lead to some insight as to how knowledge encoded in DNA might be arranged. Humans are very good at patterns and I suspect patterns may turn out to be key.
With all Best Wishes
Rick Standley”
Well…Only an interesting example might shows the tiny (whatever) bit of validity of that “language’s might survive human life in DNA storage” idea.
Before the example, I declare that, I haven’t read the wide range of content of this site, just browsing, and read some, like this.
The example: Our mother tongue, even if existing in a superb web of meaningful connections, and unlike many other languages (a studied only eights of them…) are still clearly acceptable for contemporary readers if You read a 500 years old book or any text. Without any modification or rule to apply. Sadly, the Latin language tried to latinize it and later the German also tried to replace it with tricky modifications, as now the English does (via coumputer and science and posh language, phrases, etc…) without success. I guess the english will finally achieve it, even if the language is now archived. This language is the magyar, Hungarian. The reason why I telling this, is that no matter how “superlogiacal” this language seems like – which You use in the daily bases to understand more things of Our world – there is one part of the grammar, where there is no rules apply. Not even alternative scientist or theory builders tries to explain that, it just unknown. Only a few percent of native speakers knows exactly the right way in the written grammar (if You have to write letter “j” or “ly” in spell), and trust Me, We use to have pods around Us in the elementary school when we had a grammar exam, because all the others, who had no idea what to write were inspired to check Our writing, and copy the right way. They just had no clue. And once again, there is no rule for that, and we are a very few whom knowing it, and on top of it, we know it all the cases, missing none! I thought about this so many times, since the first class of elementary school, and always ending up some sort of DNA storage or something like that, in brain. Have to mention another one thing: this language has a completely regulated grammar, like others, this is the only example, where there’s no rules or clues or patterns apply. One more thing to add, it is a very old language compared to the others in use today, unfortunately the rest of its letter, books, artifact are damaged in central Europe over the always busy historical events, like wars and revolutions and stuff. Thanks God, We kept it over hard times, and archived and digitalized it in the last few decades. Many other languages or its keepers weren’t that strong in human cultures around the globe. Hope it helps to understand more or stronger. I think this unknown example absolutely prove it. Good reading, thanks!
I for one would like a summary of the above article and how we might discern how it applies to my life – it was just to complicated tho I am fascinated by the subject.
Susan, the take away on a personal note is that we are most likely at the pinnacle of a recurring cycle of technology in which we begin to understand our connection with each other and nature. This will lead us to uploading our consciousness into machines at first, but those machines, will become biotech and further will eventually simply be nature.
Therefore on spiritual level it is likely that the spirits we sometimes commune with via plant teachers are actually the consciousness of ancients which have already ascended into that plane in the same way we will. They exist currently inside the giant computer that is nature and just like watching a modern computer can tell you nothing of the beautiful vistas and adventures unfolding inside the whirring drives and electrical signals, watching the machinery of their computer (which we call nature) reveals nothing particular about the world they experience.
Specifically it also means that there is a mechanical explanation for ideas like past life regression that a hardened materialist can follow. It means there is a slower more methodical path into spirituality that the more closed minded can follow.
But the article itself is just about the technicals of the process of information storage in DNA specifically with all the various technologies and design principles which point toward this process having happened before.
excellent.. can someone please forward this to gmo team ? 😉
Great piece 🙂
David Zindell proposed this idea in the 1980s in “The Broken God” (an absolute must-read, just do it) with the central theme of the book that an advanced ancient race had coded their memories into human DNA.
Zindell’s book did not gain traction as the astonishing ideas were too ‘out there’ for a mainstream audience but he was ahead of the curve; we are starting to catch up to some of it now…
Biotech is at the cusp of simply being another computer system that uses the same programming methodology we use in electronic computers: http://phys.org/news/2016-03-language-cells.html